Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A14 F35 I1 R1
|
74 |
857.2 |
37938082 |
99.4% |
37710453 |
136.3 |
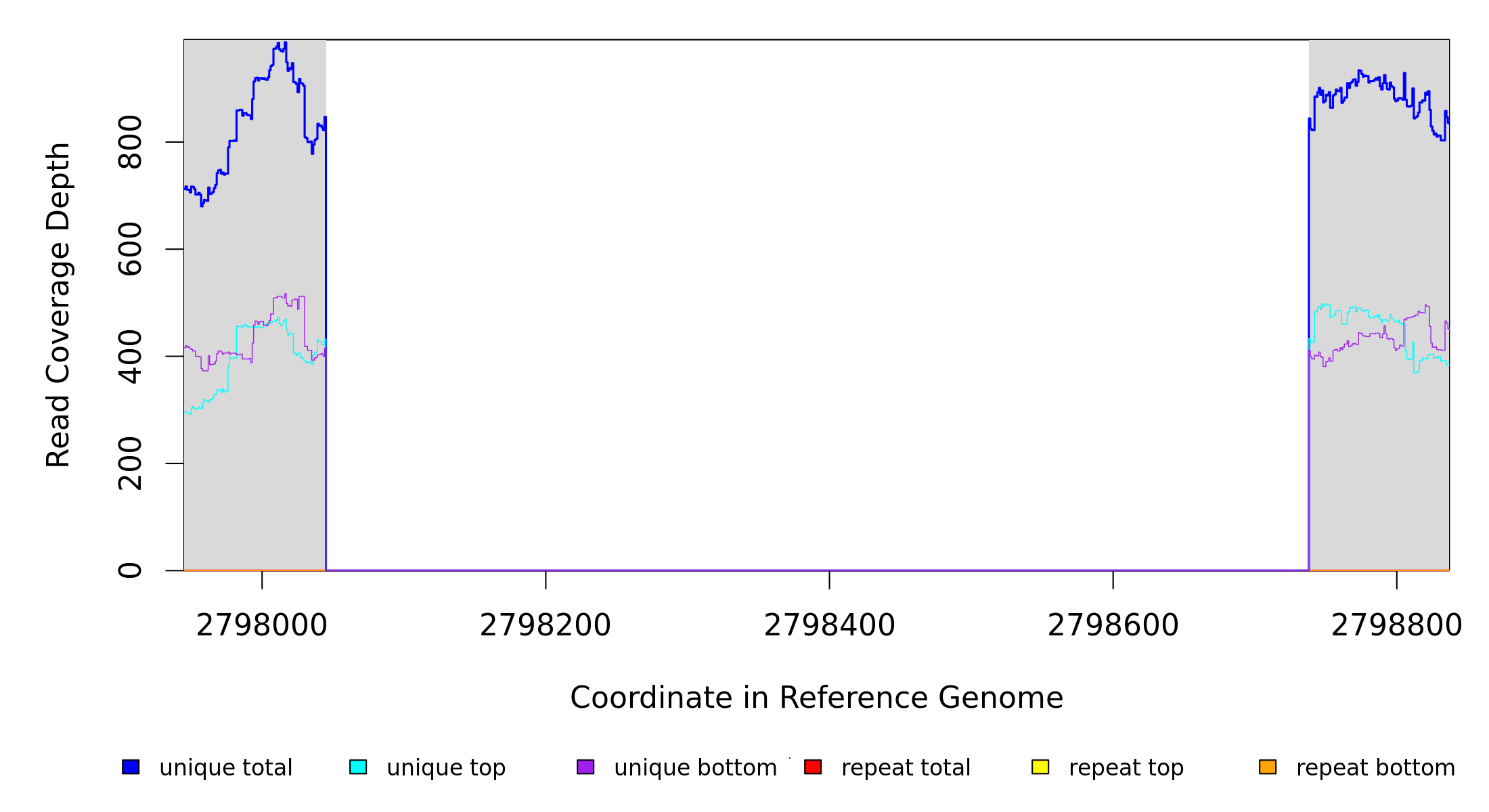
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
AE015451_phaM1 |
2,798,045 |
Δ693 bp |
coding (1‑693/693 nt) |
endX CDS (extracellular DNA endonuclease) ← |
|
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
AE015451_phaM1 |
2798045 |
2798737 |
693 |
847 [0] |
[0] 843 |
endX CDS (extracellular DNA endonuclease) |
|
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
AE015451_phaM1 |
= 2798044 | 0 (0.000) | 842 (0.980) |
121/270 |
0.3 |
100% |
intergenic (+3/+1) |
PP_2450 CDS (conserved protein of unknown function)/endX CDS (extracellular DNA endonuclease) |
–/– |
| ? | AE015451_phaM1 |
2798738 = |
0 (0.000) | intergenic (‑1/+6) |
endX CDS (extracellular DNA endonuclease)/PP_2452 CDS (conserved protein of unknown function) |
–/– |
GATK/CNVnator alignment
N/A