Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A1 F1 I1 R1
|
521 |
238.2 |
12299412 |
96.0% |
11807435 |
93.0 |
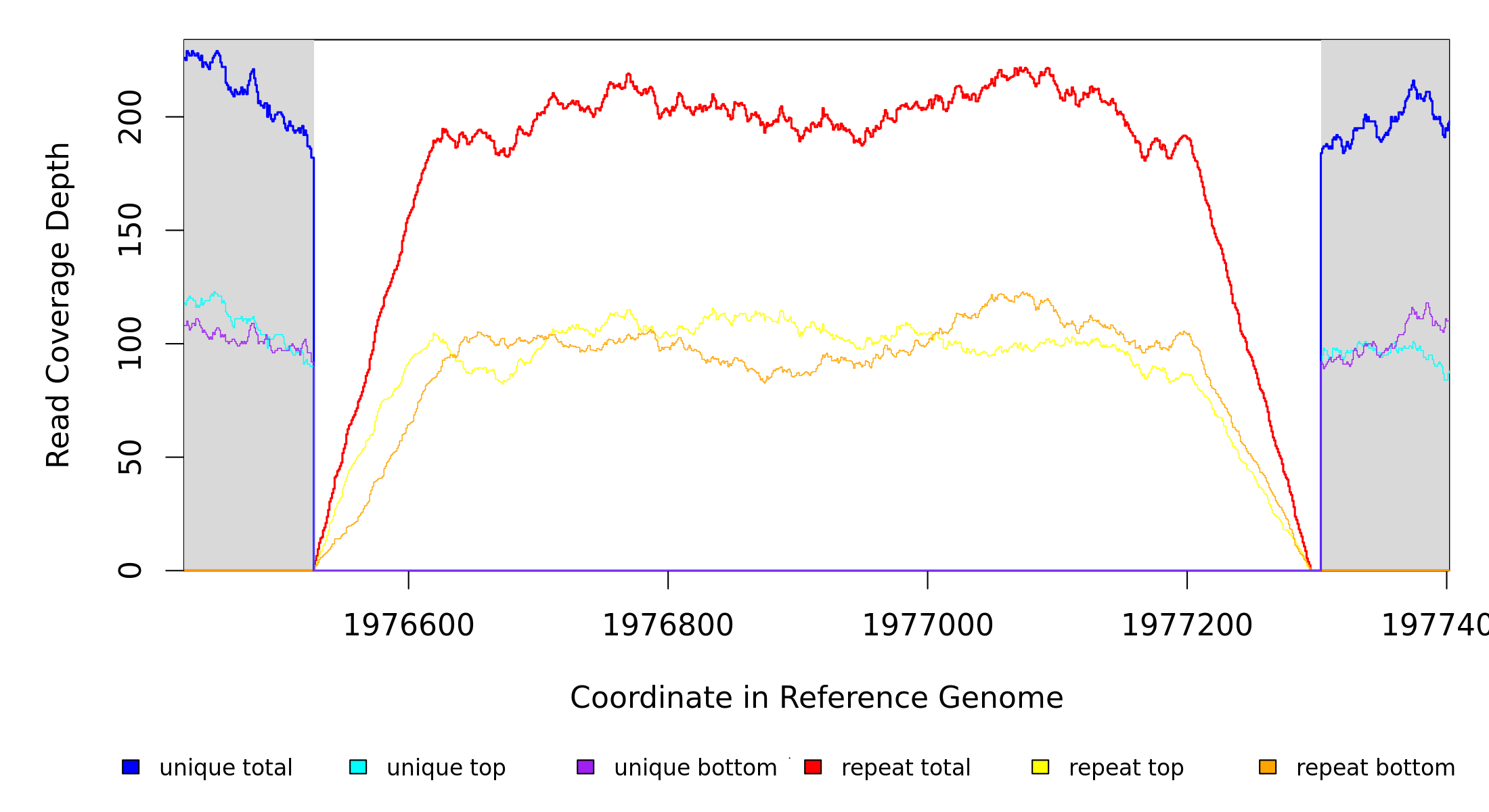
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
NC_000913 |
1,976,527 |
Δ776 bp |
|
insB5–insA5 |
insB5, insA5 |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
NC_000913 |
1976527–1977294 |
1977302 |
9–776 |
182 [0] |
[0] 184 |
insB5–insA5 |
insB5,insA5 |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
NC_000913 |
= 1976526 | 0 (0.000) | 164 (0.770) |
95/168 |
0.9 |
100% |
intergenic (‑305/+16) |
b1892/insB5 |
FlhD/IS1 protein InsB |
| ? | NC_000913 |
1977303 = |
0 (0.000) | intergenic (‑64/‑474) |
insA5/yecG |
IS1 protein InsA/putative regulator |
GATK/CNVnator alignment
N/A