Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A6 F11 I4 R1
|
73 |
908.3 |
39027734 |
99.3% |
38754539 |
136.2 |
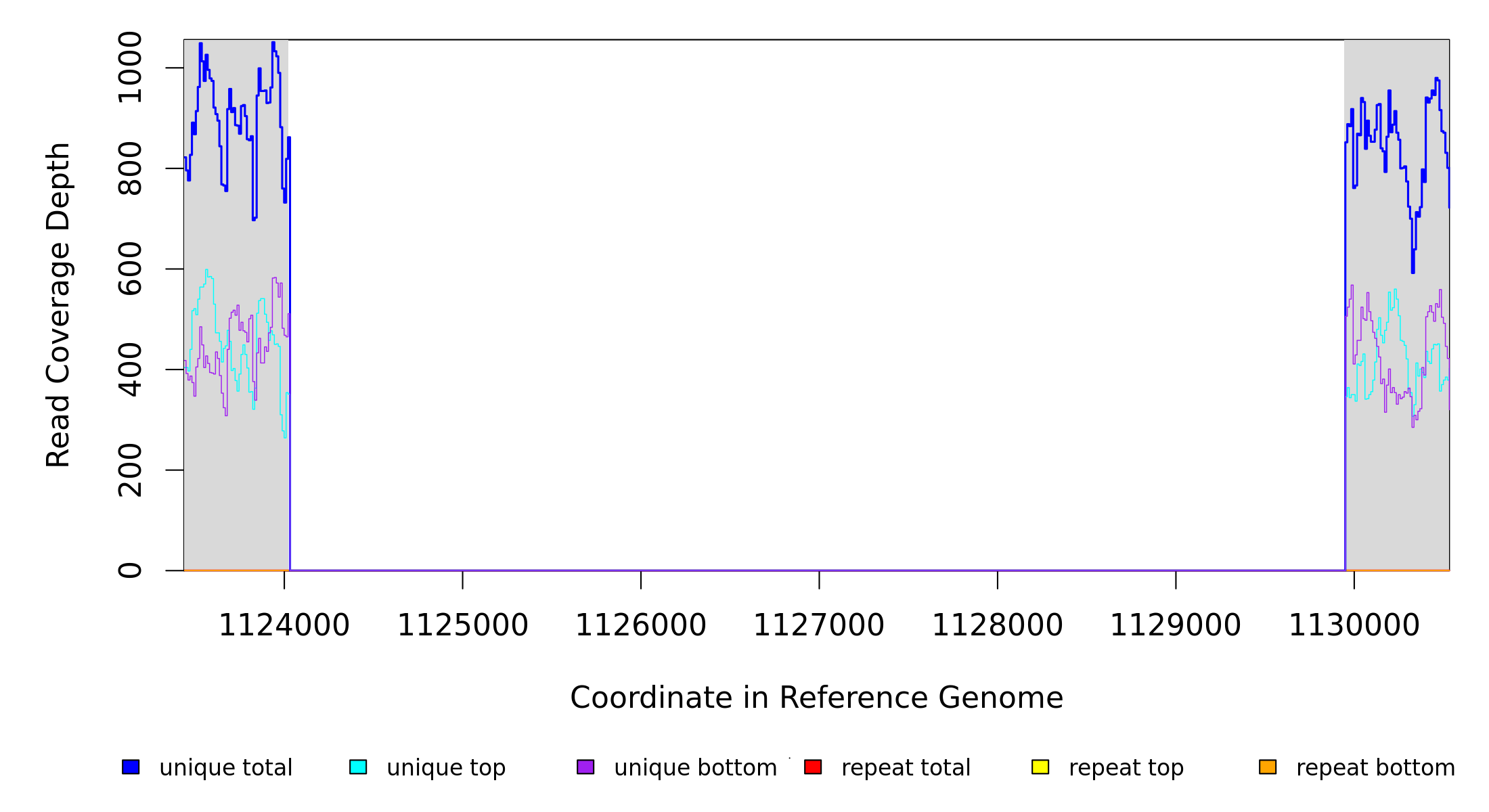
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
AE015451_phaM1 |
1,124,024 |
Δ5,919 bp |
|
[gcvT‑I CDS (aminomethyltransferase)]–[gcvH‑I CDS (glycine cleavage system H protein 1)] |
[gcvT‑I CDS (aminomethyltransferase)], tdcG‑II CDS (L‑serine dehydratase), gcvP‑I CDS (glycine dehydrogenase), [gcvH‑I CDS (glycine cleavage system H protein 1)] |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
AE015451_phaM1 |
1124024 |
1129942 |
5919 |
855 [0] |
[0] 859 |
[gcvT‑I CDS (aminomethyltransferase)]–[gcvH‑I CDS (glycine cleavage system H protein 1)] |
[gcvT‑I CDS (aminomethyltransferase)],tdcG‑II CDS (L‑serine dehydratase),gcvP‑I CDS (glycine dehydrogenase),[gcvH‑I CDS (glycine cleavage system H protein 1)] |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
AE015451_phaM1 |
= 1124023 | 0 (0.000) | 855 (0.940) |
133/270 |
0.1 |
100% |
coding (1120/1122 nt) |
gcvT‑I CDS (aminomethyltransferase) |
|
| ? | AE015451_phaM1 |
1129943 = |
0 (0.000) | coding (3/384 nt) |
gcvH‑I CDS (glycine cleavage system H protein 1) |
|
GATK/CNVnator alignment
N/A