Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A1 F1 I1 R1
|
154 |
26.6 |
2069536 |
90.2% |
1866721 |
108.9 |
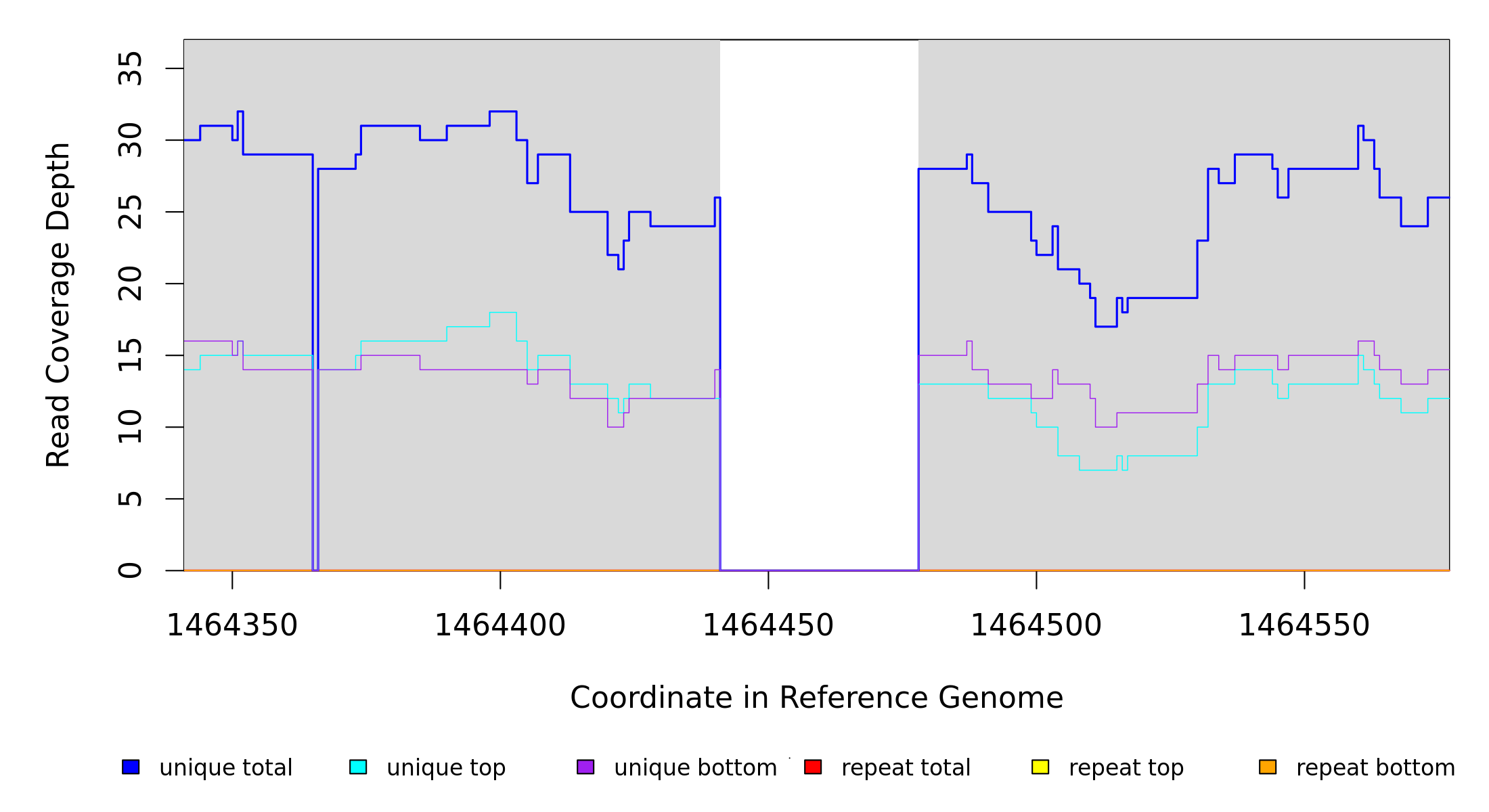
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
AM260479 |
1,464,441 |
Δ37 bp |
coding (125‑161/552 nt) |
h16_A1354 → |
conserved hypothetical protein |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
AM260479 |
1464441 |
1464477 |
37 |
26 [0] |
[0] 28 |
h16_A1354 |
conserved hypothetical protein |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
AM260479 |
= 1464440 | 0 (0.000) | 26 (0.990) |
23/216 |
0.2 |
100% |
coding (124/552 nt) |
h16_A1354 |
conserved hypothetical protein |
| ? | AM260479 |
1464478 = |
0 (0.000) | coding (162/552 nt) |
h16_A1354 |
conserved hypothetical protein |
GATK/CNVnator alignment
BRESEQ :: bam2aln output
GACAGGAGATGCCATGCACGCCTGTGACCTGTGTCATCAGCTGGTGGGCGAACCGTCCAGCAGTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAACTGCTGGTGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACGGCATTGGGTGAGCCGCCGAATATGTGGCGAATGGGCCCGCGCCCGGTCGAC > AM260479/1464304‑1464567
|
gagatgtgtataagagacaGCCTGTGACCTGTGTCATCAGCTGGTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTG < GWNJ‑0478:712:GW2002102894th:2:2101:7469:63797/131‑1 (MQ=60)
ATGCACGCCTGTGACCTGTGTCATCAGCTGGTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATG > GWNJ‑0478:712:GW2002102894th:2:1106:1634:8122/1‑150 (MQ=60)
CCTGTGACCTGTGTCATCAGCTGGTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTACC > GWNJ‑0478:712:GW2002102894th:2:2205:18402:48831/1‑147 (MQ=60)
acggcatacgagatgtagctccgtctcgtgggctcggagatgtgtataagagacagCCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGC < GWNJ‑0478:712:GW2002102894th:2:2113:18604:62230/94‑1 (MQ=60)
GACCTGTGTCATCAGCTGGTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACG > GWNJ‑0478:712:GW2002102894th:2:1108:7005:61705/1‑150 (MQ=60)
ttttttttttcaagcagaagacggcatacgagatgtagctccgtctcgtgggctcggagatgtgtataagagacagTGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACGgc < GWNJ‑0478:712:GW2002102894th:2:1210:6210:86554/73‑3 (MQ=38)
GTCATCAGCTGGTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGTCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTctgtctcttatacacatc > GWNJ‑0478:712:GW2002102894th:2:2214:3563:38645/1‑132 (MQ=60)
CATCAGCTGGTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACGgcattgggt < GWNJ‑0478:712:GW2002102894th:2:2208:11971:45971/150‑10 (MQ=60)
CATCAGCTGGTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACGgcattgggt < GWNJ‑0478:712:GW2002102894th:2:2212:11302:74673/150‑10 (MQ=60)
GTGGGCGAACCGTCCAGC‑GTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTctgtctcttatacacatctccgagcccacgagacggagctacat > GWNJ‑0478:712:GW2002102894th:2:1109:14344:50570/1‑106 (MQ=60)
GTGCAAGGTCGACAGCCGCTGGCGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCctgtctcttatacacatctccgagcccacgagacggagctacatctcgtatgccgtcttctgcttgaaaaaaaaaaaaaaaaaaaagaaaa > GWNJ‑0478:712:GW2002102894th:2:1207:18615:34373/1‑58 (MQ=16)
|
GACAGGAGATGCCATGCACGCCTGTGACCTGTGTCATCAGCTGGTGGGCGAACCGTCCAGCAGTGCCGCCACACGAGCACCTGGCTGCCTCTGGCATTGCGGCCATGAGCAACCGGTGCAAGGTCGACAACTGCTGGTGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACGCTGCACGGGCTGCGGCGCCTGGATGTACCAGAGCACGGCATTGGGTGAGCCGCCGAATATGTGGCGAATGGGCCCGCGCCCGGTCGAC > AM260479/1464304‑1464567
|
| Alignment Legend |
|---|
Aligned base mismatch/match (shaded by quality score): ATCG/ATCG < 0 ≤ ATCG/ATCG < 35 ≤ ATCG/ATCG < 38 ≤ ATCG/ATCG < 40 ≤ ATCG/ATCG |
Unaligned base: atcg Masked matching base: atcg Alignment gap: ‑ Deleted base: ‑ |