Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A9 F92 I6 R1
|
18 |
65.3 |
1472746 |
69.2% |
1019140 |
300.6 |
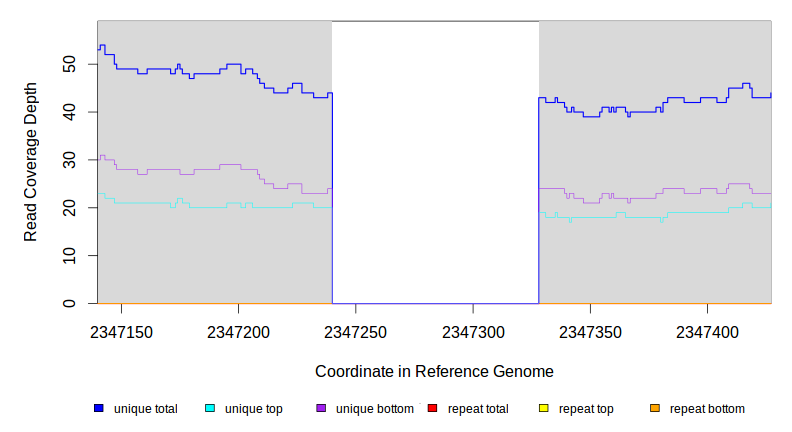
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
NC_000913 |
2,347,240 |
Δ88 bp |
intergenic (+90/‑57) |
nrdA → / → nrdB |
ribonucleoside‑diphosphate reductase 1, alpha subunit/ribonucleoside‑diphosphate reductase 1, beta subunit, ferritin‑like protein |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
NC_000913 |
2347240 |
2347327 |
88 |
44 [0] |
[0] 43 |
nrdA/nrdB |
ribonucleoside‑diphosphate reductase 1, alpha subunit/ribonucleoside‑diphosphate reductase 1, beta subunit, ferritin‑like protein |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
NC_000913 |
= 2347239 | 0 (0.000) | 39 (0.690) |
38/522 |
0.5 |
100% |
noncoding (80/203 nt) |
REP164 (repetitive extragenic palindromic) element; contains 5 REP sequences |
REP164 (repetitive extragenic palindromic) element; contains 5 REP sequences |
| ? | NC_000913 |
2347328 = |
0 (0.000) | noncoding (169/203 nt) |
REP164 (repetitive extragenic palindromic) element; contains 5 REP sequences |
REP164 (repetitive extragenic palindromic) element; contains 5 REP sequences |
GATK/CNVnator alignment
N/A