Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A16 F258 I0 R1
|
432 |
103.0 |
5191697 |
92.7% |
4812703 |
99.8 |
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
freq |
annotation |
gene |
description |
| MC JC |
NC_000913 |
3,582,206 |
Δ1,222 bp |
100% |
IS1‑mediated |
[yhhZ]–yrhA |
[yhhZ], yrhA |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
NC_000913 |
3582206 |
3584146–3583427 |
1222–1941 |
113 [3] |
[52] 53 |
[yhhZ]–[insB1] |
[yhhZ], yrhA, insA, [insB1] |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
NC_000913 |
= 3582205 | 3 (0.030) | 99 (1.080) |
70/178 |
NT |
97.1% |
coding (343/1179 nt) |
yhhZ |
putative Hcp1 family polymorphic toxin protein; putative colicin‑like DNase/tRNase activity |
| ? | NC_000913 |
3583428 = |
NA (NA) | noncoding (1/768 nt) |
IS1 |
repeat region |
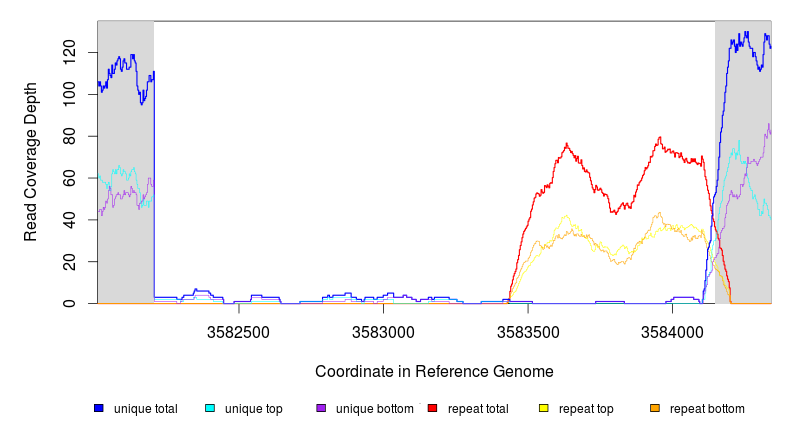
GATK/CNVnator alignment
N/A