Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A2 F17 I0 R1
|
43 |
55.2 |
1410130 |
81.3% |
1146435 |
215.5 |
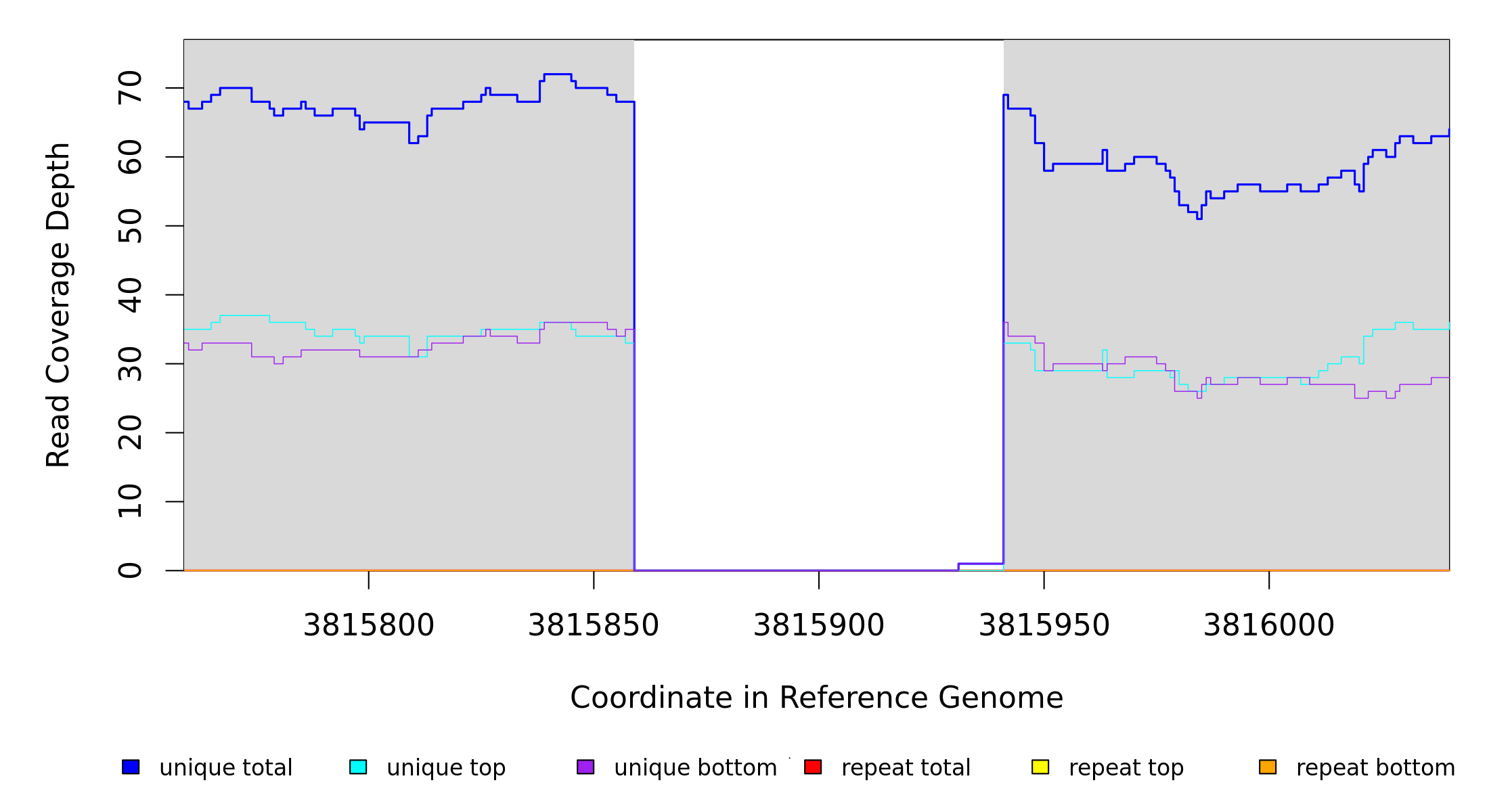
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
freq |
annotation |
gene |
description |
| MC JC |
NC_000913 |
3,815,859 |
Δ82 bp |
100% |
|
[rph]–[rph] |
[rph], [rph] |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
NC_000913 |
3815859 |
3815940 |
82 |
68 [0] |
[1] 69 |
[rph]–[rph] |
[rph],[rph] |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
NC_000913 |
= 3815858 | 0 (0.000) | 65 (1.380) |
46/388 |
NT |
100% |
pseudogene (24/48 nt) |
rph |
ribonuclease PH (defective);enzyme; Degradation of RNA; RNase PH |
| ? | NC_000913 |
3815941 = |
0 (0.000) | pseudogene (609/669 nt) |
rph |
ribonuclease PH (defective);enzyme; Degradation of RNA; RNase PH |
GATK/CNVnator alignment
BRESEQ :: bam2aln output
GTATTAAACAGCCCGGCGTTGAAGAAATAGGGGCTTTTGCGCCCGGATTTCAGCGTAAACTCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGCCGCCTTCTGCGTCGCTACAATGGATTCGATTCCCCTCGGGCCAGAGCCAACAAGATGAGTAGCTCTTCATGGGTGAACGGCTCGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAATTCCGACAGAAACTGCGGCTACCATCCCTTTCAT > NC_000913/3815644‑3816087
|
GTATTAAACAGCCCGGCGTTGAAGAAATAGGGGCTTTTGCGCCCGGATTTCAGCGTAAACTCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTTATCTTACTTTTCTAACGACAAAAAAAAGGCGATTCATCAGTCGCCTTAAAAATCAGTTTGCAAGCGCAGCCTTCTGCGgtcccctgcacttcaatgatgcgcccgtctttggt > M01186:129:000000000‑A5M8V:1:2114:16920:8586/1‑216 (MQ=60)
TCAGCGTAAACGCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCACGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCC < M01186:129:000000000‑A5M8V:1:2106:10348:10617/251‑1 (MQ=60)
CGTAAACTCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGAT < M01186:129:000000000‑A5M8V:1:1109:14784:19610/251‑1 (MQ=60)
CCGCCAAACTTTAACACCTGCTTGCTAAGCCCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAAC < M01186:129:000000000‑A5M8V:1:1110:26051:12216/251‑1 (MQ=60)
TCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTATTCTGTCATCACTCCGTTCATGTCGGTCTCTGCGGCCGAGTCTTCACCGTATTCCAGCTCGCAAACCCCTTCGCCGTTCACAATTCCGCCAGAAACTCCGG > M01186:129:000000000‑A5M8V:1:1112:29347:13115/1‑250 (MQ=60)
CAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACGTTCATGTCGGTCTCTCCGGCAGAGCATTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAATTCCGACAGAAACTGCGGCA > M01186:129:000000000‑A5M8V:1:1107:6401:23468/1‑251 (MQ=60)
GATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGCTCGCAAACCGCTTCTCCGTTCACAATTCCGACAGAAACTTCGGCTACCATCCCTTTCAT > M01186:129:000000000‑A5M8V:1:2107:21686:25535/1‑250 (MQ=60)
|
GTATTAAACAGCCCGGCGTTGAAGAAATAGGGGCTTTTGCGCCCGGATTTCAGCGTAAACTCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGCCGCCTTCTGCGTCGCTACAATGGATTCGATTCCCCTCGGGCCAGAGCCAACAAGATGAGTAGCTCTTCATGGGTGAACGGCTCGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAATTCCGACAGAAACTGCGGCTACCATCCCTTTCAT > NC_000913/3815644‑3816087
|
| Alignment Legend |
|---|
Aligned base mismatch/match (shaded by quality score): ATCG/ATCG < 0 ≤ ATCG/ATCG < 12 ≤ ATCG/ATCG < 14 ≤ ATCG/ATCG < 30 ≤ ATCG/ATCG < 39 ≤ ATCG/ATCG |
Unaligned base: atcg Masked matching base: atcg Alignment gap: ‑ Deleted base: ‑ |