Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A3 F28 I0 R1
|
51 |
55.9 |
1320120 |
86.3% |
1139263 |
218.0 |
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
freq |
annotation |
gene |
description |
| MC JC |
NC_000913 |
3,815,859 |
Δ82 bp |
100% |
|
[rph]–[rph] |
[rph], [rph] |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
NC_000913 |
3815859 |
3815940 |
82 |
77 [0] |
[0] 77 |
[rph]–[rph] |
[rph],[rph] |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
NC_000913 |
= 3815858 | 0 (0.000) | 74 (1.550) |
53/394 |
NT |
100% |
pseudogene (24/48 nt) |
rph |
ribonuclease PH (defective);enzyme; Degradation of RNA; RNase PH |
| ? | NC_000913 |
3815941 = |
0 (0.000) | pseudogene (609/669 nt) |
rph |
ribonuclease PH (defective);enzyme; Degradation of RNA; RNase PH |
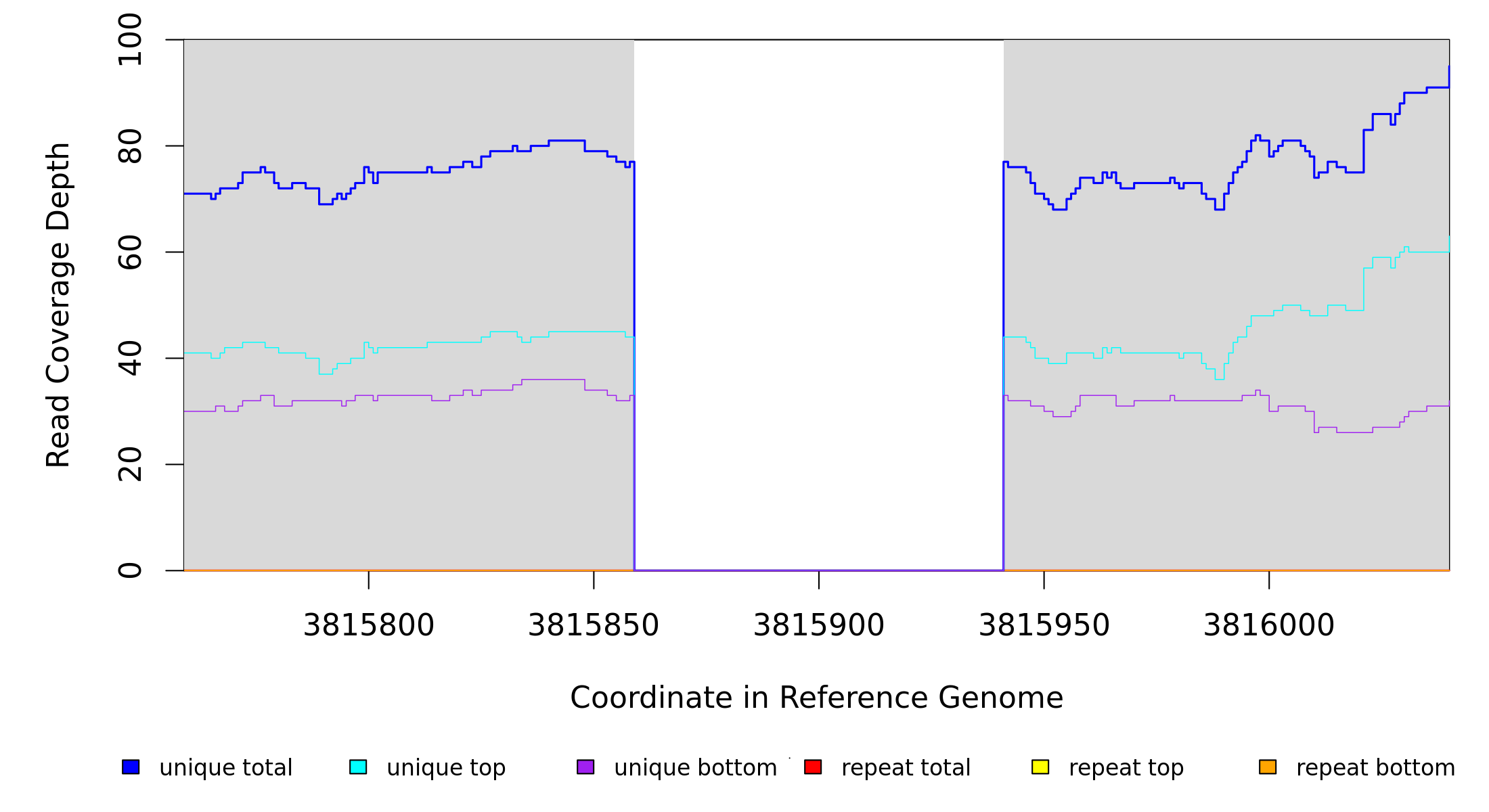
GATK/CNVnator alignment
BRESEQ :: bam2aln output
CGTAAACTCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGCCGCCTTCTGCGTCGCTACAATGGATTCGATTCCCCTCGGGCCAGAGCCAACAAGATGAGTAGCTCTTCATGGGTGAACGGCTCGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAATTCCGACAGAAACTGCGGCTACCATCCCTTTCATCG > NC_000913/3815697‑3816089
|
CGTAAACTCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAG < M01186:129:000000000‑A5M8V:1:1104:13918:20951/249‑1 (MQ=60)
CTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCCCAAACCACTTCC > M01186:129:000000000‑A5M8V:1:1101:13682:23042/1‑250 (MQ=60)
TGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAAT > M01186:129:000000000‑A5M8V:1:2109:12700:18932/1‑251 (MQ=60)
ATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCCGAATTCCGCCCGAACCTGCG > M01186:129:000000000‑A5M8V:1:2105:16476:3073/1‑251 (MQ=60)
TTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTACCGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAATTCCGACAGAAACTGAG > M01186:129:000000000‑A5M8V:1:2103:15394:16607/1‑251 (MQ=60)
GATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTCCTCCGTTCACAATTACGCCAGAAACTGCGCGTGACATCCCTTTCAT > M01186:129:000000000‑A5M8V:1:2107:15081:20014/1‑250 (MQ=60)
ATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGTCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGC‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑‑CGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAATTCCGACAGAAACTGCGGCTACCATCCCTTTCATCG > M01186:129:000000000‑A5M8V:1:2109:20329:22359/1‑251 (MQ=60)
|
CGTAAACTCGCCAAACTTTAACACCTGCTTGCTAAGCGCAAATTCAATAAACTGGCGCTGATATGGTTTCATGCCTTCGCTCCTCATCTTACTTTTCTACAGACAAAAAAAAGGCGACTCATCAGTCGCCTTAAAAATCAGTTTGCCAGCGCCGCCTTCTGCGTCGCTACAATGGATTCGATTCCCCTCGGGCCAGAGCCAACAAGATGAGTAGCTCTTCATGGGTGAACGGCTCGCCTTCTGCCGTCCCCTGCACTTCAATGATGCGCCCGTCTTCGGTCATCACTA‑CGTTCATGTCGGTCTCTGCGGCAGAGTCTTCAACGTATTCCAGATCGCAAACCGCTTCGCCGTTCACAATTCCGACAGAAACTGCGGCTACCATCCCTTTCATCG > NC_000913/3815697‑3816089
|
| Alignment Legend |
|---|
Aligned base mismatch/match (shaded by quality score): ATCG/ATCG < 0 ≤ ATCG/ATCG < 12 ≤ ATCG/ATCG < 14 ≤ ATCG/ATCG < 33 ≤ ATCG/ATCG < 39 ≤ ATCG/ATCG |
Unaligned base: atcg Masked matching base: atcg Alignment gap: ‑ Deleted base: ‑ |