Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A2 F1 I1 R1
|
404 |
265.1 |
10478538 |
96.8% |
10143224 |
150.7 |
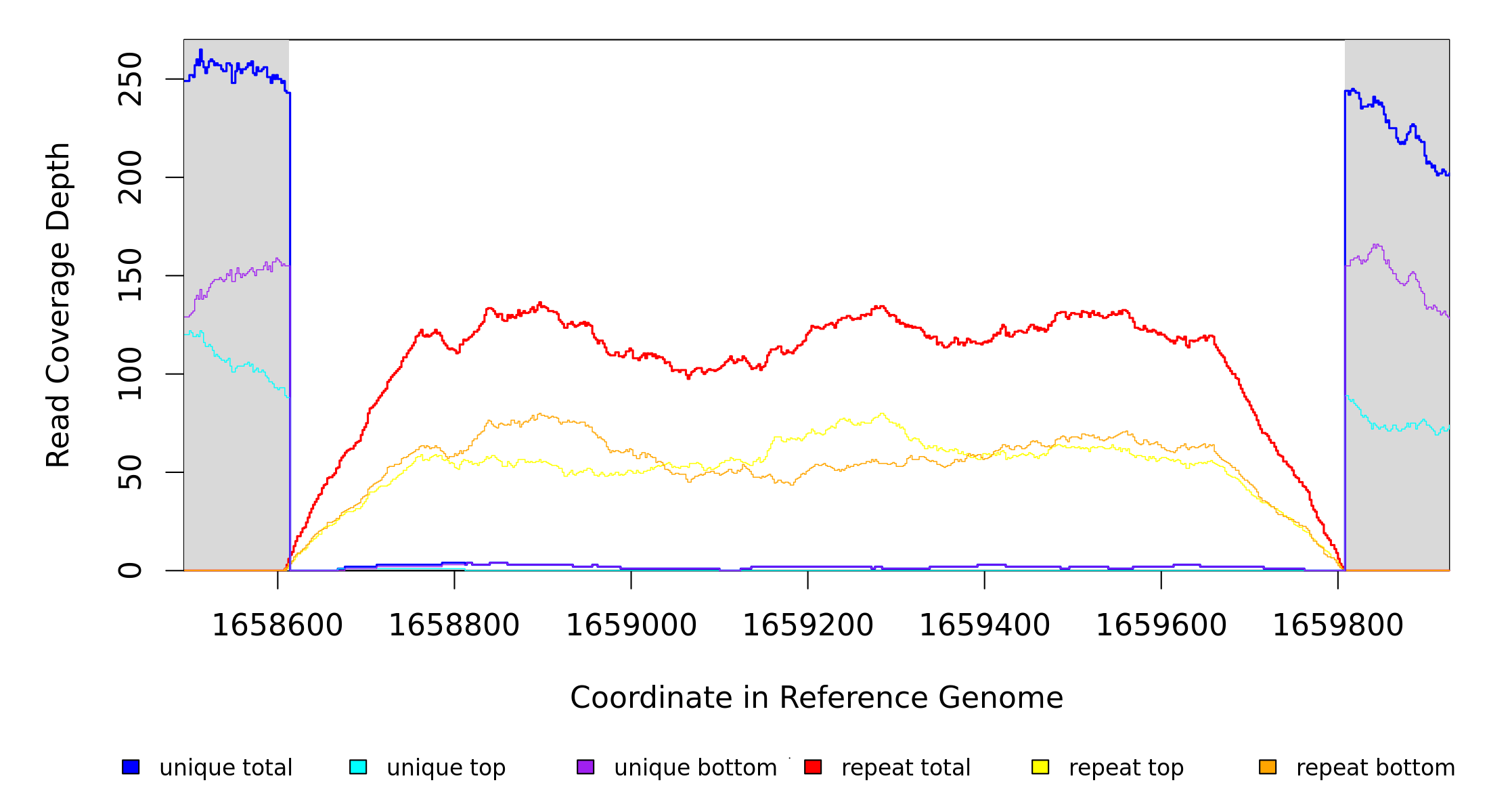
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
CP009974 |
1,658,613 |
Δ1,195 bp |
|
RPPX_07425 |
RPPX_07425 |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
CP009974 |
1658613–1659806 |
1659807 |
2–1195 |
243 [0] |
[0] 244 |
RPPX_07425 |
RPPX_07425 |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
CP009974 |
= 1658612 | 0 (0.000) | 238 (0.940) |
156/290 |
0.3 |
100% |
intergenic (‑17/‑64) |
RPPX_07420/RPPX_07425 |
acyl‑CoA dehydrogenase/transposase |
| ? | CP009974 |
1659808 = |
0 (0.000) | intergenic (+152/+271) |
RPPX_07425/RPPX_07430 |
transposase/pyrroloquinoline quinone biosynthesis protein PqqE |
GATK/CNVnator alignment
N/A