Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate | Predicted Mutations | Mean Coverage | Total Reads | Percent Mapped | Mapped Reads | Average Read Length |
|---|---|---|---|---|---|---|
| A25 F999 I0 R1 | 111 | 24.8 | 2365438 | 97.5% | 2306302 | 48.6 |
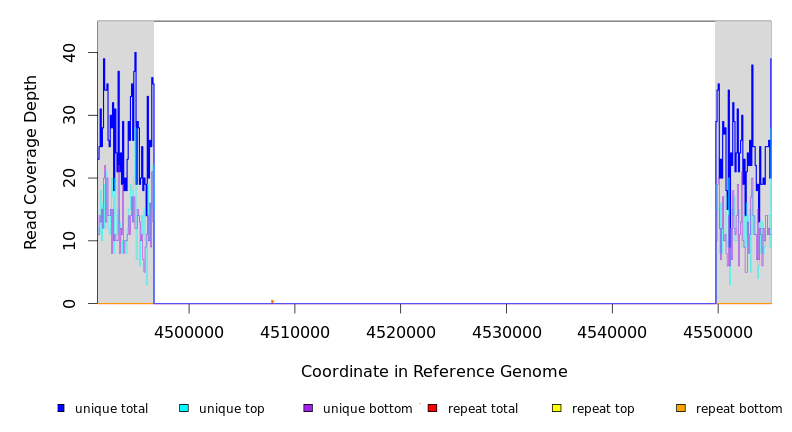
Breseq alignment
| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | NC_000913 | 4,496,676 | Δ53,036 bp | intB–fimH | 54 genes intB, insC‑6, insD‑6, yjgX, yjgZ, insG, yjhB, yjhC, ythA, yjhD, yjhE, insO, insI‑3, insO, insO, yjhV, fecE, fecD, fecC, fecB, fecA, fecR, fecI, insA‑7, yjhU, yjhF, yjhG, yjhH, yjhI, sgcR, sgcE, sgcA, ryjB, sgcQ, sgcC, sgcB, sgcX, yjhP, yjhQ, topAI, yjhZ, yjhR, nanS, nanM, nanC, fimB, fimE, fimA, fimI, fimC, fimD, fimF, fimG, fimH |
|
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
| * | * | ÷ | NC_000913 | 4496676 | 4549711 | 53036 | 26 [0] | [0] 26 | intB–fimH | intB,insC‑6,insD‑6,yjgX,yjgZ,insG,yjhB,yjhC,ythA,yjhD,yjhE,insO,insI‑3,insO,insO,yjhV,fecE,fecD,fecC,fecB,fecA,fecR,fecI,insA‑7,yjhU,yjhF,yjhG,yjhH,yjhI,sgcR,sgcE,sgcA,ryjB,sgcQ,sgcC,sgcB,sgcX,yjhP,yjhQ,topAI,yjhZ,yjhR,nanS,nanM,nanC,fimB,fimE,fimA,fimI,fimC,fimD,fimF,fimG,fimH |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | NC_000913 | = 4496675 | 0 (0.000) | 25 (0.840) | 24/94 | 0.1 | 100% | intergenic (+186/‑75) | leuX/intB | tRNA‑Leu/KpLE2 phage‑like element; putative integrase |
| ? | NC_000913 | 4549712 = | 0 (0.000) | intergenic (+2/+241) | fimH/gntP | type 1 fimbriae D‑mannose specific adhesin/fructuronate transporter | |||||