Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate | Predicted Mutations | Mean Coverage | Total Reads | Percent Mapped | Mapped Reads | Average Read Length |
|---|---|---|---|---|---|---|
| A26 F999 I0 R1 | 111 | 30.9 | 2867207 | 97.9% | 2806995 | 48.6 |
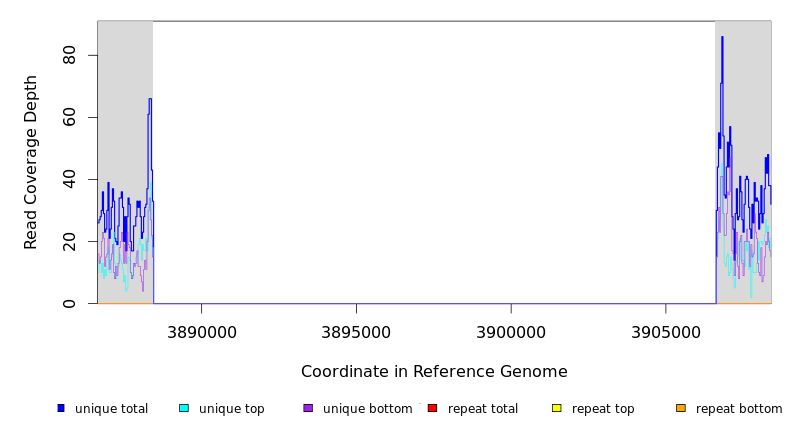
Breseq alignment
| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | NC_000913 | 3,888,432 | Δ18,167 bp | tnaC–bglG | 17 genes tnaC, tnaA, tnaB, mdtL, yidZ, yieE, yieF, adeP, yieH, cbrB, cbrC, yieK, yieL, bglH, bglB, bglF, bglG |
|
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
| * | * | ÷ | NC_000913 | 3888432 | 3906598 | 18167 | 30 [0] | [0] 31 | tnaC–bglG | tnaC,tnaA,tnaB,mdtL,yidZ,yieE,yieF,adeP,yieH,cbrB,cbrC,yieK,yieL,bglH,bglB,bglF,bglG |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | NC_000913 | = 3888431 | 0 (0.000) | 29 (0.800) | 26/94 | 0.1 | 100% | intergenic (+239/‑4) | mnmE/tnaC | 5‑carboxymethylaminomethyluridine‑tRNA synthase GTPase subunit/tnaAB operon leader peptide |
| ? | NC_000913 | 3906599 = | 0 (0.000) | intergenic (‑32/+254) | bglG/phoU | transcriptional antiterminator BglG/negative regulator of the pho regulon | |||||