Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate | Predicted Mutations | Mean Coverage | Total Reads | Percent Mapped | Mapped Reads | Average Read Length |
|---|---|---|---|---|---|---|
| A10 F999 I0 R1 | 75 | 85.3 | 6433650 | 96.2% | 6189171 | 49.6 |
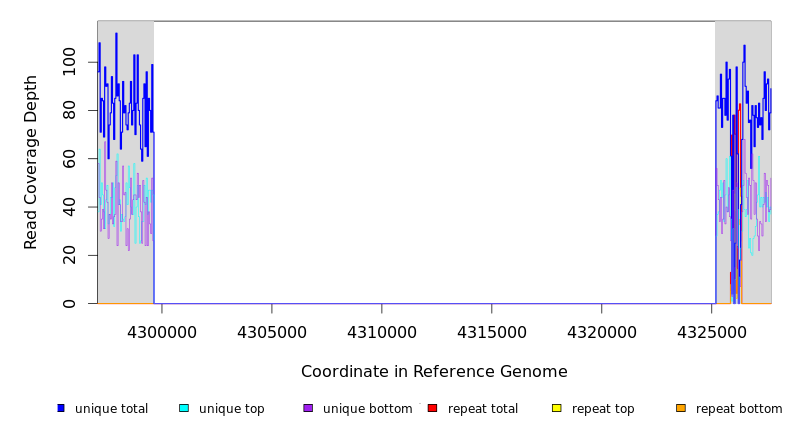
Breseq alignment
| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | NC_000913 | 4,299,628 | Δ25,538 bp | [mdtP]–[phnC] | 27 genes [mdtP], mdtO, mdtN, ytcA, yjcS, alsK, alsE, alsC, alsA, alsB, alsR, rpiB, yjdP, phnP, phnO, phnN, phnM, phnL, phnK, phnJ, phnI, phnH, phnG, phnF, phnE, phnD, [phnC] |
|
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
| * | * | ÷ | NC_000913 | 4299628 | 4325165 | 25538 | 71 [0] | [0] 70 | [mdtP]–[phnC] | [mdtP],mdtO,mdtN,ytcA,yjcS,alsK,alsE,alsC,alsA,alsB,alsR,rpiB,yjdP,phnP,phnO,phnN,phnM,phnL,phnK,phnJ,phnI,phnH,phnG,phnF,phnE,phnD,[phnC] |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | NC_000913 | = 4299627 | 0 (0.000) | 66 (0.790) | 44/98 | 0.1 | 100% | coding (1404/1467 nt) | mdtP | putative multidrug efflux pump outer membrane channel |
| ? | NC_000913 | 4325166 = | 0 (0.000) | intergenic (‑1/+132) | phnC/yjdN | phosphonate ABC transporter ATP binding subunit/conserved protein YjdN | |||||