Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate | Predicted Mutations | Mean Coverage | Total Reads | Percent Mapped | Mapped Reads | Average Read Length |
|---|---|---|---|---|---|---|
| A10 F999 I0 R1 | 75 | 85.3 | 6433650 | 96.2% | 6189171 | 49.6 |
Breseq alignment
| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | NC_000913 | 4,555,491 | Δ41,523 bp | [yjiC]–lgoD | 37 genes [yjiC], ytiC, ytiD, idlP, iraD, hypT, iadA, yjiG, yjiH, kptA, yjiJ, yjiK, ytiA, yjiL, yjiM, yjiN, mdtM, rpnD, yjiR, yjiS, yjiT, yjiV, mcrC, mcrB, symE, symR, hsdS, hsdM, hsdR, mrr, yjiA, yjiX, btsT, tsr, lgoT, lgoR, lgoD |
|
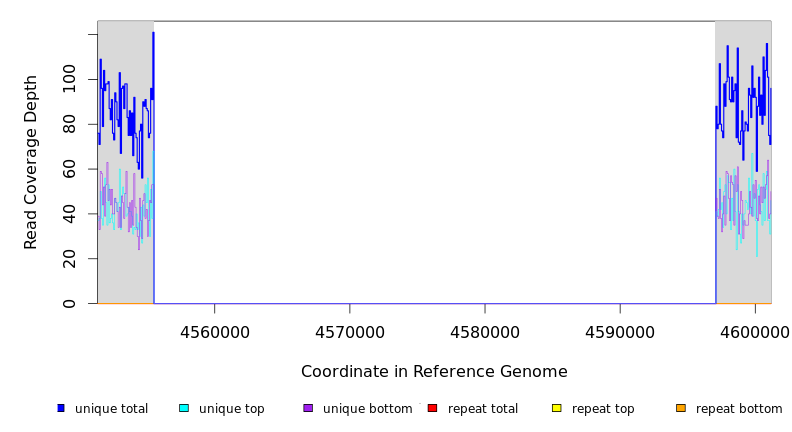
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
| * | * | ÷ | NC_000913 | 4555491 | 4597013 | 41523 | 79 [0] | [0] 78 | [yjiC]–lgoD | [yjiC],ytiC,ytiD,idlP,iraD,hypT,iadA,yjiG,yjiH,kptA,yjiJ,yjiK,ytiA,yjiL,yjiM,yjiN,mdtM,rpnD,yjiR,yjiS,yjiT,yjiV,mcrC,mcrB,symE,symR,hsdS,hsdM,hsdR,mrr,yjiA,yjiX,btsT,tsr,lgoT,lgoR,lgoD |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | NC_000913 | = 4555490 | 0 (0.000) | 72 (0.890) | 49/96 | 0.0 | 100% | coding (831/831 nt) | yjiC | uncharacterized protein YjiC |
| ? | NC_000913 | 4597014 = | 0 (0.000) | intergenic (+2/+136) | lgoD/opgB | L‑galactonate oxidoreductase/phosphoglycerol transferase I | |||||