Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate | Predicted Mutations | Mean Coverage | Total Reads | Percent Mapped | Mapped Reads | Average Read Length |
|---|---|---|---|---|---|---|
| A8 F60 I1 R1 | 13 | 156.7 | 5387192 | 99.4% | 5354868 | 139.8 |
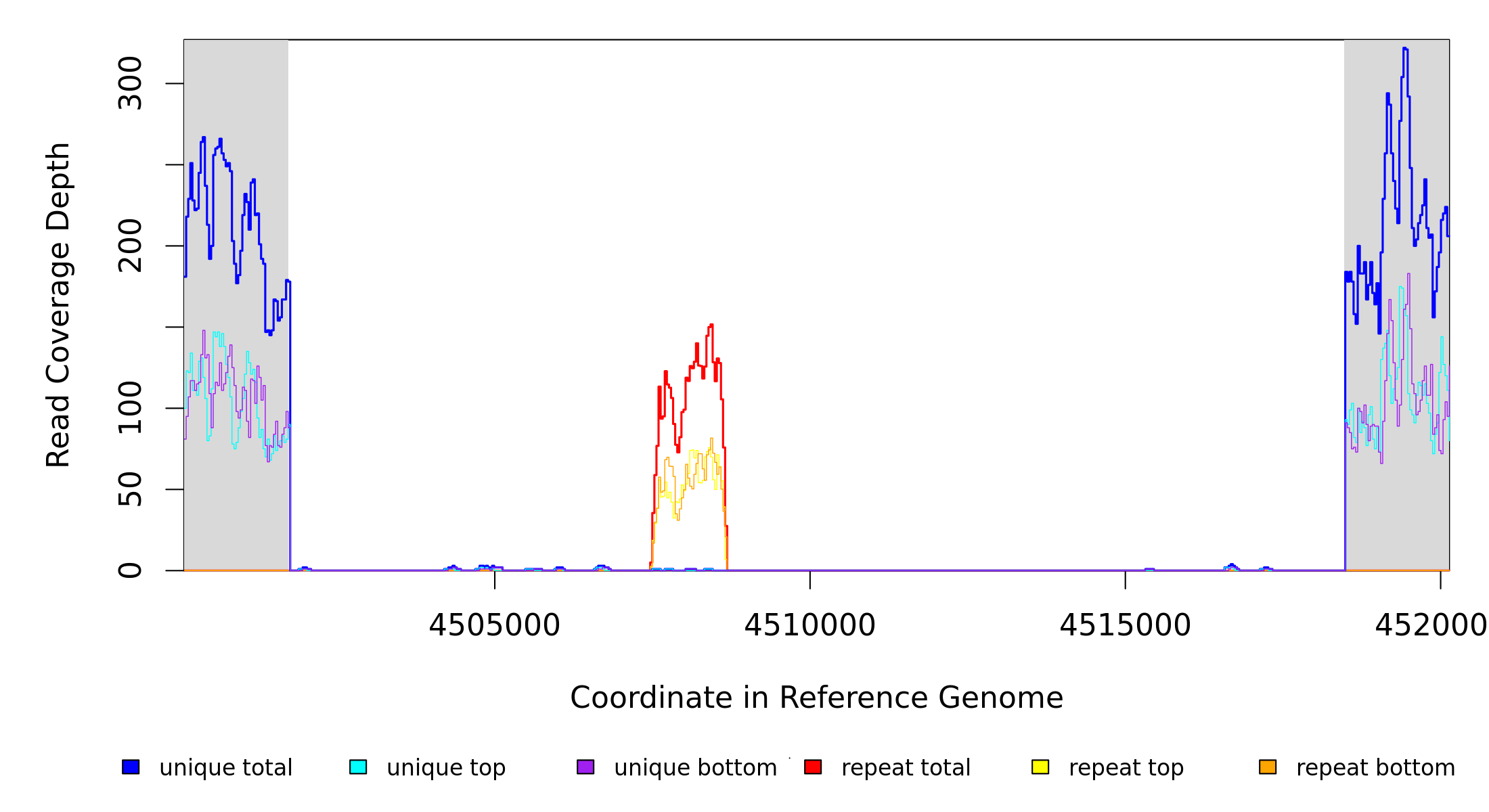
Breseq alignment
| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | NC_000913 | 4,501,724 | Δ16,748 bp | IS1‑mediated | insG–fecI | 18 genes insG, yjhB, yjhC, ythA, yjhD, yjhE, insO, insI‑3, insO, insO, yjhV, fecE, fecD, fecC, fecB, fecA, fecR, fecI |
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
| * | * | ÷ | NC_000913 | 4501724 | 4518471 | 16748 | 178 [0] | [0] 179 | insG–fecI | insG,yjhB,yjhC,ythA,yjhD,yjhE,insO,insI‑3,insO,insO,yjhV,fecE,fecD,fecC,fecB,fecA,fecR,fecI |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | NC_000913 | = 4501723 | 0 (0.000) | 178 (1.110) | 96/278 | 0.1 | 100% | intergenic (+134/+380) | yjgZ/insG | KpLE2 phage‑like element; uncharacterized protein YjgZ/KpLE2 phage‑like element; IS4 putative transposase |
| ? | NC_000913 | 4518472 = | NA (NA) | noncoding (1/768 nt) | IS1 | repeat region | |||||