Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate | Predicted Mutations | Mean Coverage | Total Reads | Percent Mapped | Mapped Reads | Average Read Length |
|---|---|---|---|---|---|---|
| A52 F1 I1 R1 | 19 | 66.0 | 4263516 | 97.2% | 4144137 | 76.0 |
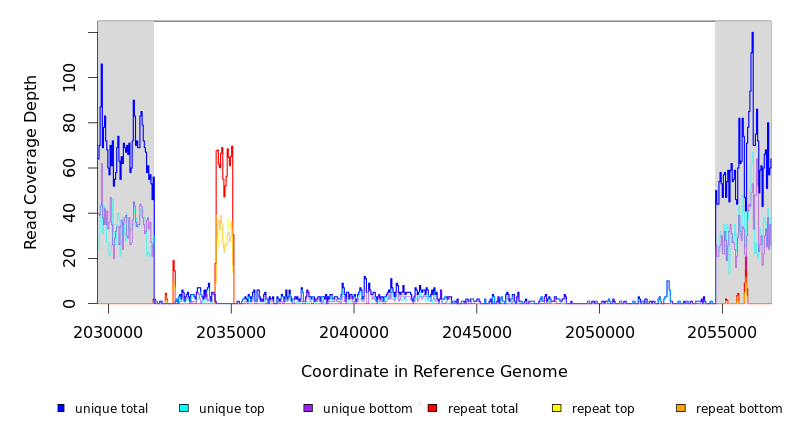
Breseq alignment
| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | REL606 | 2,031,850 | Δ22,863 bp | [manB]–wcaJ | 22 genes [manB], manC, insB‑14, insA‑14, wbbD, wbbC, wzy, wbbB, wbbA, vioB, vioA, wzx, rmlC, rfbA, rfbD, rfbB, galF, wcaM, wcaL, wcaK, wzxC, wcaJ |
|
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
| * | * | ÷ | REL606 | 2031850–2031888 | 2054712 | 22825–22863 | 55 [0] | [1] 56 | [manB]–wcaJ | [manB],manC,insB‑14,insA‑14,wbbD,wbbC,wzy,wbbB,wbbA,vioB,vioA,wzx,rmlC,rfbA,rfbD,rfbB,galF,wcaM,wcaL,wcaK,wzxC,wcaJ |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | REL606 | = 2031849 | 0 (0.000) | 55 (0.830) | 41/148 | 0.6 | 99.1% | coding (949/1365 nt) | manB | Phosphomannomutase |
| ? | REL606 | 2054713 = | 1 (0.020) | intergenic (‑42/+13) | wcaJ/cpsG | predicted UDP‑glucose lipid carrier transferase/phosphomannomutase | |||||