Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A1 F2 I194 R1
|
197 |
32.2 |
1845014 |
95.2% |
1756453 |
85.4 |
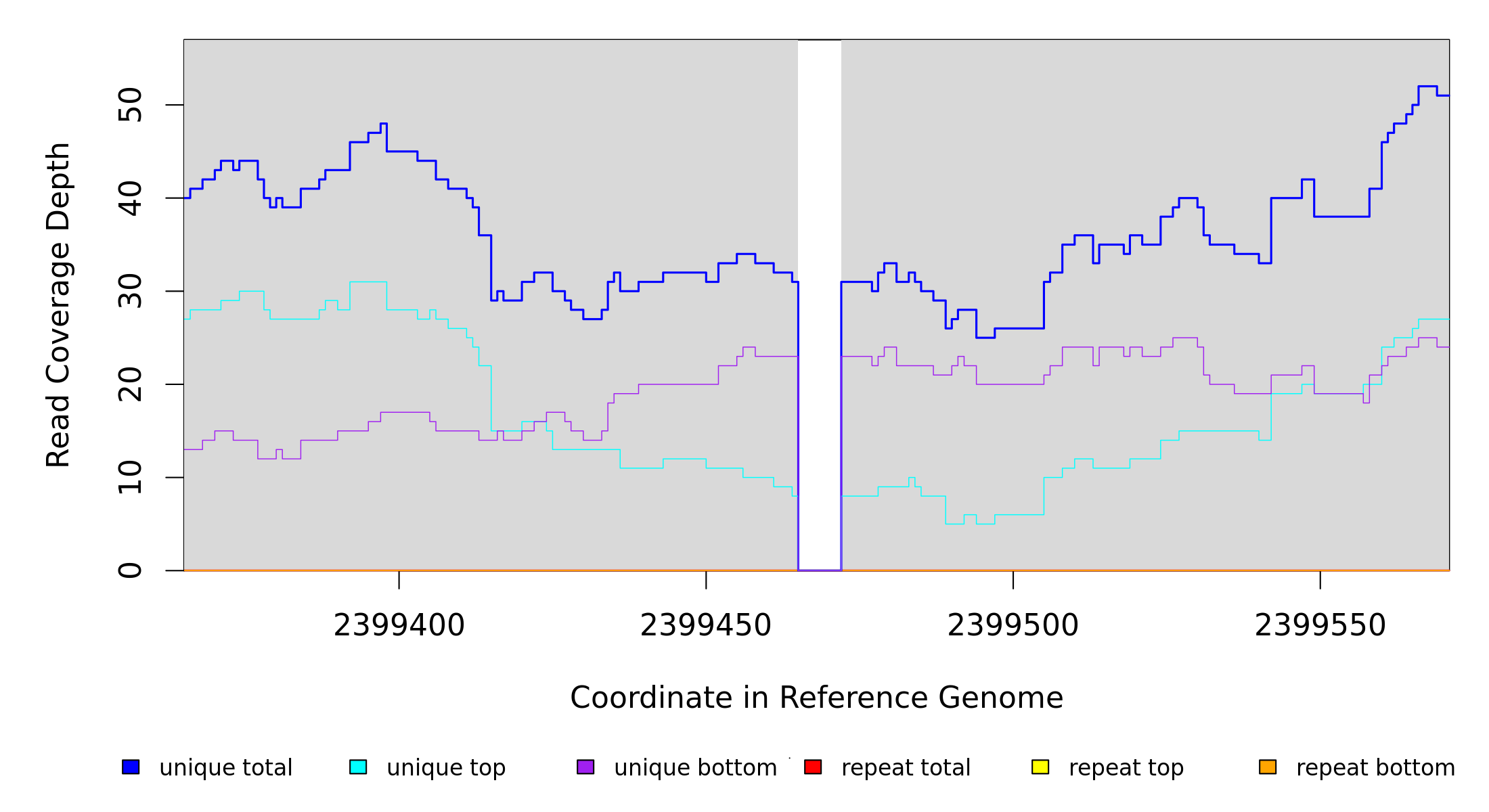
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
NZ_CP009273 |
2,399,465 |
Δ7 bp |
coding (650‑656/939 nt) |
lrhA ← |
transcriptional regulator LrhA |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
NZ_CP009273 |
2399465 |
2399471 |
7 |
31 [0] |
[0] 31 |
lrhA |
transcriptional regulator LrhA |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
NZ_CP009273 |
= 2399464 | 0 (0.000) | 31 (0.970) |
23/168 |
0.2 |
100% |
coding (657/939 nt) |
lrhA |
transcriptional regulator LrhA |
| ? | NZ_CP009273 |
2399472 = |
0 (0.000) | coding (649/939 nt) |
lrhA |
transcriptional regulator LrhA |
GATK/CNVnator alignment
BRESEQ :: bam2aln output
ACGCAGGTCCGGGCTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAGGCGACATAAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATATCGCGAAACGGGCTAGGATCATCCAGCAATACAAGAGGGAT > NZ_CP009273/2399377‑2399562
|
ACGCAGGTCCGGGCTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCC > SRR3722072.470243/1‑100 (MQ=60)
GNTCCGGGCTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGA > SRR3722072.23265/1‑100 (MQ=60)
GGTCCGGGCTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGA > SRR3722072.342595/1‑100 (MQ=60)
GGTCCGGGCTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGA > SRR3722072.896568/1‑100 (MQ=60)
GCTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTG < SRR3722072.611735/100‑1 (MQ=60)
CTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGC < SRR3722072.150816/100‑1 (MQ=60)
CATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGCTTTAT > SRR3722072.902521/1‑100 (MQ=60)
CACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATAT < SRR3722072.156821/100‑1 (MQ=60)
TTACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATATCGCGAAACGGGCTAGGA > SRR3722072.248349/1‑100 (MQ=60)
CACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATATCGCGAAACGGGCTAGGAT < SRR3722072.338227/100‑1 (MQ=60)
CACTGCCGCACGAACGGCCGGAAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATATCGCGAAACGGGCTAGGAT < SRR3722072.748102/100‑1 (MQ=60)
AAGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATATCGCGAAACGGGCTAGGATCATCCAGCAATACAAGAGGGA < SRR3722072.904714/100‑1 (MQ=60)
AGCGTCGAG‑‑‑‑‑‑‑AAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATATCGCGAAACGGGCTAGGATCATCCAGCAATACAAGAGGGAT < SRR3722072.574621/100‑1 (MQ=60)
|
ACGCAGGTCCGGGCTCATCATCTCAACCGGCCTTGCCGTCACGCCAAGACCGGCTTTCACTGCCGCACGAACGGCCGGAAGCGTCGAGGCGACATAAGCCAGTCGCCATGGAATATCTGCTTTATTAAGCGTCGCCAGCACCATATCGCGAAACGGGCTAGGATCATCCAGCAATACAAGAGGGAT > NZ_CP009273/2399377‑2399562
|
| Alignment Legend |
|---|
Aligned base mismatch/match (shaded by quality score): ATCG/ATCG < 0 ≤ ATCG/ATCG < 23 ≤ ATCG/ATCG < 30 ≤ ATCG/ATCG < 35 ≤ ATCG/ATCG < 41 ≤ ATCG/ATCG |
Unaligned base: atcg Masked matching base: atcg Alignment gap: ‑ Deleted base: ‑ |