Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate |
Predicted Mutations |
Mean Coverage |
Total Reads |
Percent Mapped |
Mapped Reads |
Average Read Length |
|
A3 F108 I2 R1
|
13 |
72.2 |
2506112 |
94.4% |
2365769 |
142.8 |
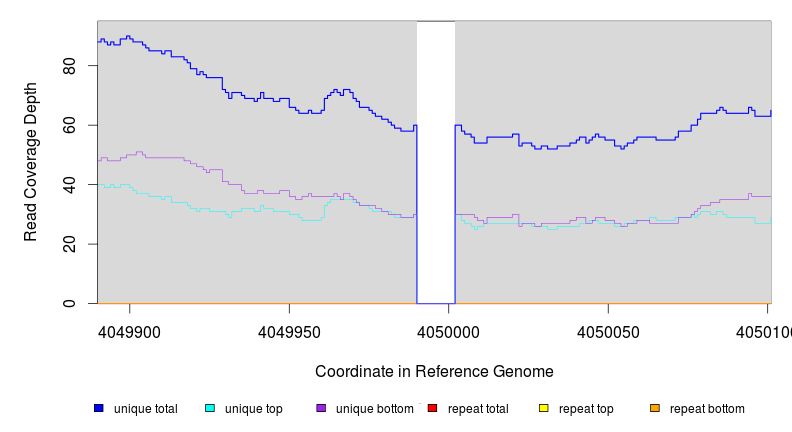
Breseq alignment
BRESEQ :: Evidence
|
| evidence |
seq id |
position |
mutation |
annotation |
gene |
description |
| MC JC |
NC_000913 |
4,049,990 |
Δ12 bp |
noncoding (93‑104/109 nt) |
spf → |
Spot 42 sRNA antisense regulator of galK translation, Hfq‑dependent |
|
| | | | seq id |
start |
end |
size |
←reads |
reads→ |
gene |
description |
|---|
| * |
* |
÷ |
NC_000913 |
4049990 |
4050001 |
12 |
60 [0] |
[0] 60 |
spf |
Spot 42 sRNA antisense regulator of galK translation, Hfq‑dependent |
|
| |
seq id |
position |
reads (cov) |
reads (cov) |
score |
skew |
freq |
annotation |
gene |
product |
| * |
? |
NC_000913 |
= 4049989 | 0 (0.000) | 58 (0.810) |
45/282 |
0.6 |
100% |
noncoding (92/109 nt) |
spf |
Spot 42 sRNA antisense regulator of galK translation, Hfq‑dependent |
| ? | NC_000913 |
4050002 = |
NA (NA) | noncoding (105/109 nt) |
spf |
Spot 42 sRNA antisense regulator of galK translation, Hfq‑dependent |
GATK/CNVnator alignment
N/A