Sample Resequencing Stats
Note: The mutation counts shown below represent unfiltered mutation sets.
| ALE, Flask, Isolate | Predicted Mutations | Mean Coverage | Total Reads | Percent Mapped | Mapped Reads | Average Read Length |
|---|---|---|---|---|---|---|
| A18 F1 I1 R1 | 18 | 50.6 | 3235256 | 96.4% | 3118786 | 76.0 |
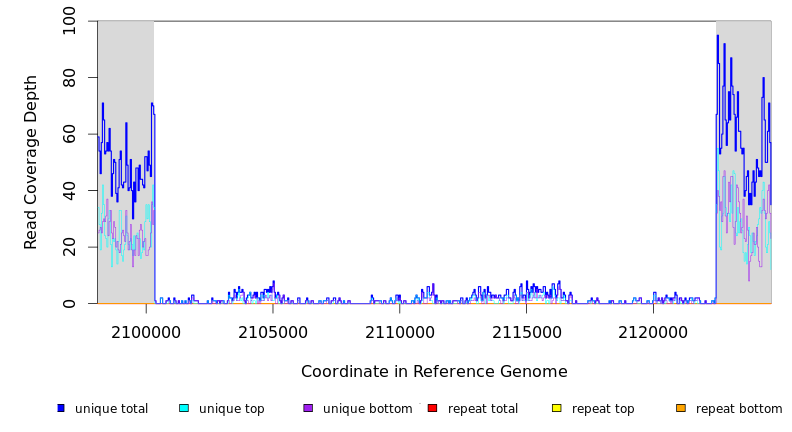
Breseq alignment
| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | REL606 | 2,100,308 | Δ22,146 bp | ogrK–[ECB_02013] | 27 genes ogrK, yegZ, ECB_01989, ECB_01990, ECB_01991, ECB_01992, ECB_01993, ECB_01994, ECB_01995, ECB_01996, ECB_01997, ECB_01998, ECB_01999, ECB_02000, ECB_02001, ECB_02002, ECB_02003, ECB_02004, ECB_02005, ECB_02006, ECB_02007, ECB_02008, ECB_02009, ECB_02010, ECB_02011, ECB_02012, [ECB_02013] |
|
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
| * | * | ÷ | REL606 | 2100308 | 2122453 | 22146 | 68 [1] | [3] 69 | ogrK–[ECB_02013] | ogrK,yegZ,ECB_01989,ECB_01990,ECB_01991,ECB_01992,ECB_01993,ECB_01994,ECB_01995,ECB_01996,ECB_01997,ECB_01998,ECB_01999,ECB_02000,ECB_02001,ECB_02002,ECB_02003,ECB_02004,ECB_02005,ECB_02006,ECB_02007,ECB_02008,ECB_02009,ECB_02010,ECB_02011,ECB_02012,[ECB_02013] |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | REL606 | = 2100307 | 1 (0.020) | 49 (1.410) | 38/104 | 0.0 | 97.3% | intergenic (+171/+102) | yegQ/ogrK | predicted peptidase/phage DNA‑binding transcriptional regulator |
| ? | REL606 | 2122454 = | 2 (0.060) | coding (172/216 nt) | ECB_02013 | conserved hypothetical protein; putative exported protein | |||||